CS81 Adaptive Robotics¶
Lab 5: Applying Novelty Search¶
Due the Monday after Fall Break (Oct. 23)¶
Lab Goals¶
- Combine Novelty Search with NEAT to solve the coverage problem.
- Compare Novelty + NEAT results to the NEAT results you got in Lab 3.
Getting Started¶
Use the github server to get the starting point notebooks for this lab.
Introduction¶
In this lab you will combine your implementation of novelty search with NEAT and apply it to the coverage task. The goal of novelty search is to find unique behaviors, rather than optimizing a particular objective function. The pictures below show some example behaviors that were in the archive at the end of a single novelty search run with a population of 100 that was evolved for just 5 generations. Note that many of these behaviors are not very good at coverage, however they are quite different from one another.
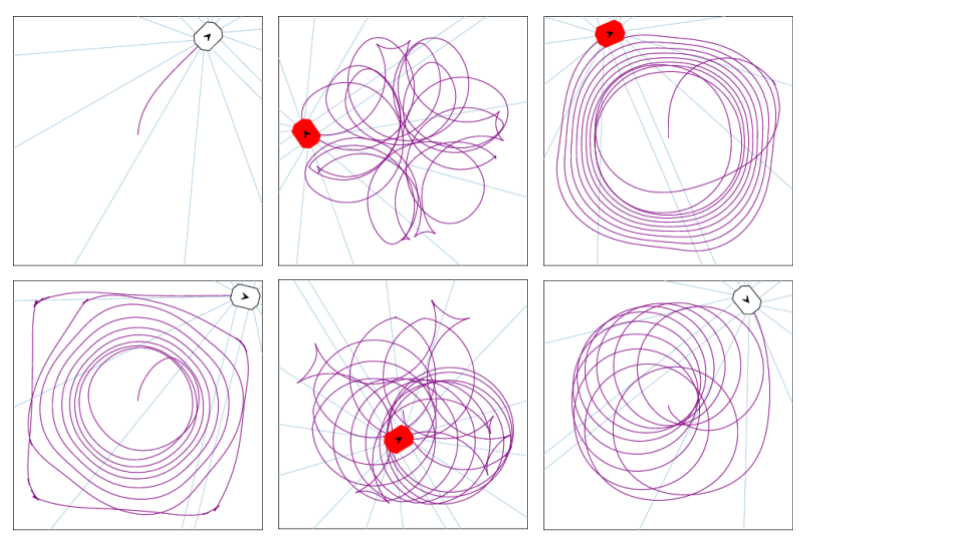
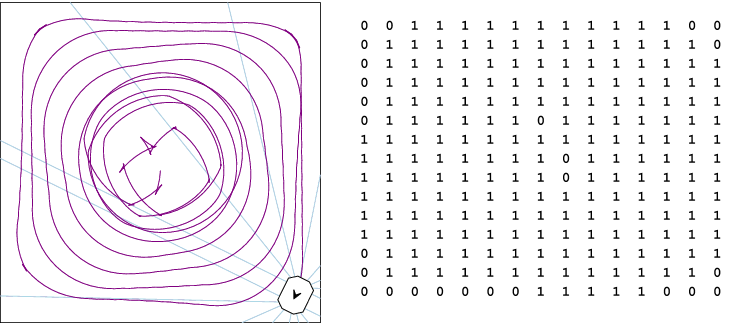
Despite the fact that novelty search is not optimizing an objective function, it is still able to find successful networks that perform coverage quite well, as demonstrated in the pictures shown below of both the trail and grid of the best network found. This network achieves 88.45% coverage, and only misses a few grid locations in the center, as well as some locations in the corners. This network had:
- 4 inputs: Representing whether the agent is stalled, a normalized distance to the nearest wall in front, a normalized count down timer, and a constant bias of 1.0.
- 2 outputs: Representing translation and rotation amounts.
This network was found in generation 23 out of a total of 25 generations. The novelty search parameters were:
- k=15
- limit=100
- threshold=0.2
- max_dist= sqrt(2 x 14 x 14) x 225)
from jyro.simulator import *
from math import pi, floor
from random import random
from neat3 import config, population, chromosome, genome, visualize
from neat3.nn import nn_pure as nn
import pickle
import numpy as np
Update and import your novelty search notebook¶
- Copy your NoveltySearch.ipynb notebook from your lab4 directory into your lab5 directory.
- Open this notebook and add two new class variables into the constructor called self.bestScore, which you should initialize to 0, and self.bestChromos, which you should initialize to the empty list. These two variables will be used to track the best objective fitness networks found during the novelty search process.
- Comment out any test code in this notebook.
- Save it.
- Use git to add, commit, and push it.
Then you can import the notebook as shown in the next cell.
%run NoveltySearch.ipynb
Configure NEAT for novelty search¶
The maximum possible novelty score is 1.0, which occurs the first time a behavior is added to the archive.
- maximum_fitness_threshold has been set to 1.1 to ensure that every novelty run executes all generations.
- elitism has been set to 0 to encourage as much diversity as possible.
- You will need to set the number of input nodes to fit the number of sensors you used in your Lab 3 experiments.
%%file configNovelty
[phenotype]
input_nodes = 4
output_nodes = 2
max_weight = 30
min_weight = -30
feedforward = 1
nn_activation = tanh
hidden_nodes = 0
weight_stdev = 0.9
[genetic]
pop_size = 100
max_fitness_threshold = 1.1
prob_addconn = 0.1
prob_addnode = 0.05
prob_mutatebias = 0.2
bias_mutation_power = 0.5
prob_mutate_weight = 0.9
weight_mutation_power = 1.5
prob_togglelink = 0.01
elitism = 0
[genotype compatibility]
compatibility_threshold = 3.0
compatibility_change = 0.0
excess_coeficient = 1.0
disjoint_coeficient = 1.0
weight_coeficient = 0.4
[species]
species_size = 10
survival_threshold = 0.2
old_threshold = 30
youth_threshold = 10
old_penalty = 0.2
youth_boost = 1.2
max_stagnation = 15
Determine the task sensors¶
Modify the get_sensors function to use your preferred sensors for the coverage task. You should use the same sensors that you experimented with in Lab 3, so that you can compare the results you get with Novelty + NEAT vs NEAT alone.
def make_world(physics):
physics.addBox(0, 0, 4, 4, fill="white", wallcolor="black")
def make_robot():
robot = Pioneer("Pioneer", 2, 2, 0)
robot.addDevice(Pioneer16Sonars())
return robot
def get_sensors(robot, steps, i):
sonars = robot["sonar"].getData()
scaled = [min(v/5.0, 1.0) for v in sonars]
timer_down = (steps-i)/steps
inputs = [min(scaled[3:5]), robot.stall, timer_down, 1.0]
return inputs
Modify the Grid class¶
We want the Grid class to create a behavior description that we can use in novelty search. We will record, in order, the first time a grid cell is visited. For these experiments, use a grid size of 15x15.
- Modify the constructor to include a class variable called self.behavior, initialized as an empty list, where you will store the behavior of the robot.
- Modify the update method so that whenever a new grid cell is visited, its (col, row) is added to the self.behavior list.
- Add a get_behavior method that returns a list of length grid_width x grid_width that begins with the contents of the self.behavior list and is padded with (0, 0) entries to make it the correct length.
class Grid(object):
"""This class creates a grid of locations on top of a simulated world
to monitor how much of the world has been visited. Each grid location
is initally set to 0 to indicate that it is unvisited, and is updated
to 1, once it has been visited."""
def __init__(self, grid_width, world_width):
self.grid_width = grid_width
self.world_width = world_width
self.grid = []
for i in range(self.grid_width):
self.grid.append([0] * self.grid_width)
def show(self):
"""Print a representation of the grid."""
for i in range(self.grid_width):
for j in range(self.grid_width):
print("%2d" % self.grid[i][j], end=" ")
print()
print()
def update(self, x, y):
"""In the simulator, the origin is at the bottom left corner.
Adjust the row to match this configuration. Set the appropriate
grid location to 1."""
size = self.world_width/self.grid_width
col = floor(x/size)
# adjust the row so that it matches the simulator
row = self.grid_width - 1 - floor(y/size)
self.grid[row][col] = 1
def analyze_visits(self):
"""Calculate the percentage of visited cells in the grid."""
cells_visited = 0
for i in range(self.grid_width):
for j in range(self.grid_width):
if self.grid[i][j] > 0:
cells_visited += 1
percent_visited = cells_visited/self.grid_width**2
return percent_visited
Determining fitness¶
The eval_individual function is used to evaluate a particular neural network brain to determine both the objective fitness (in this case how well it controls the robot's movement such that it maximizes its coverage of the world) and the behavior description used by novelty search.
The eval_novelty_population function is used to evaluate the current population. The NEAT interface requires that this function takes a single parameter representing the population. Because we need additional variables to evaluate the population, these are designated as global variables. Read through this function and make sure you understand how NEAT and novelty search have been combined. Notice that the fitness being used by NEAT is the novelty of the brain's behavior, yet we are still tracking the objective fitness so as to find what we are ultimately searching for--good coverage performance.
The evolve function takes a NoveltySearch object along with the desired number of generations, and initiates the search.
You should not need to modify any of this code.
def eval_individual(brain, robot, sim, show_trail=False, steps=1000):
"""Returns both the coverage score AND the behavior list"""
robot.setPose(2, 2, 0)
if show_trail:
robot.useTrail = True
robot.trail = []
robot.display['trail'] = 1
grid = Grid(15, 4)
for i in range(steps):
brain.flush()
inputs = get_sensors(robot, steps, i)
output = brain.sactivate(inputs)
robot.move(*output)
x, y, a = robot.getPose()
grid.update(x, y)
sim.step()
return grid.analyze_visits(), grid.get_behavior()
def eval_novelty_population(population):
global novSearch, robot, sim
bestScoreOfGen = 0
bestChromoOfGen = None
print("Evaluating chromo", end=" ")
for i in range(len(population)):
print(i, end=" ")
chromo = population[i]
brain = nn.create_ffphenotype(chromo)
objective_fitness, behavior = eval_individual(brain, robot, sim)
if objective_fitness > bestScoreOfGen:
bestScoreOfGen = objective_fitness
bestChromoOfGen = chromo
novelty = novSearch.check_archive(behavior, chromo)
chromo.fitness = novelty
print()
if bestScoreOfGen > novSearch.bestScore:
print("!!! New best coverage behavior found %f\n" % bestScoreOfGen)
novSearch.bestScore = bestScoreOfGen
novSearch.bestChromos.append((bestChromoOfGen, bestScoreOfGen))
f = open("bestChromo%d" % (len(novSearch.bestChromos)-1), "wb")
pickle.dump(bestChromoOfGen, f)
f.close()
def evolve(novSearch, generations):
config.load("configNovelty")
chromosome.node_gene_type = genome.NodeGene
population.Population.evaluate = eval_novelty_population
pop = population.Population()
pop.epoch(generations, report=True)
visualize.plot_stats(pop.stats)
visualize.plot_species(pop.species_log)
print("\nFound behaviors with the following coverage scores:")
for i in range(len(novSearch.bestChromos)):
print(novSearch.bestChromos[i][1])
print("\nNovelty scores for 5 most recently added behaviors:")
i = 0
for saved in novSearch.archive[-5:]:
print(saved[1])
f = open("novelty%d" % (i), "wb")
pickle.dump(saved[2], f)
f.close()
i += 1
novSearch.plot_growth()
Execute the search¶
Uncomment the call to the evolve function. Begin with a small number of generations (such as 2), just to be sure that everything is working properly.
Once you have successfully completed a short run, you can try a longer run of 10 or more generations. You may need to adjust the novelty search threshold so as to most effectively use the archive for your particular sensors.
robot = make_robot()
sim = Simulator(robot, make_world)
novSearch = NoveltySearch(15, 100, 0.2, np.sqrt(2*14*14)*225)
#evolve(novSearch, 2)
Analyze the results¶
Once the search completes, it generates a number of chromo files with either a bestChromo prefix for good coverage brains or with a novelty prefix for sample brains from the archive. It will also generate the average fitness plot as well as the speciation plot. Once you complete a longer run, open the average fitness plot. Why does it look so different from the one created when you did NEAT alone?
Uncomment the call to the get_results function to generate the coverage trails and phenotypes found by the search. Open these trails and see what kinds of behaviors the search has generated. Open the phenotypes to see whether Novelty + NEAT generated networks that use hidden nodes.
def test_chromo(chromo_file):
global robot, sim
config.load('configNovelty')
chromosome.node_gene_type = genome.NodeGene
fp = open(chromo_file, "rb")
chromo = pickle.load(fp)
print(chromo)
fp.close()
visualize.draw_net(chromo, "_" + chromo_file)
brain = nn.create_ffphenotype(chromo)
print("Evaluating coverage...", end=" ")
coverage, behavior = eval_individual(brain, robot, sim, show_trail=True)
canvas = Canvas((400,400))
sim.physics.draw(canvas)
canvas.save("trail_" + chromo_file + ".svg")
print(coverage)
def get_results(novSearch):
for i in range(len(novSearch.bestChromos)):
filename = "bestChromo%d" % i
print(filename)
test_chromo(filename)
for i in range(5):
filename = "novelty%d" % i
print(filename)
test_chromo(filename)
#get_results(novSearch)
Summarize your results¶
In Lab 3, you did 10 experiments with NEAT alone to solve the coverage problem. Here you should compare those results to what you find when using Novelty + NEAT in 10 experiments, where you report the same information as before.
For deceptive tasks, we would expect Novelty + NEAT to outperform NEAT alone, but for non-deceptive tasks, we would expect NEAT to be better. Is the coverage task deceptive? We would also expect Novelty + NEAT to generate a more diverse set of behaviors than NEAT alone. Do your results match these expectations? You can refer to particular trail files to support your argument.
While you are waiting for these experiments to run, you should open up the PosterExperiments.ipynb notebook and start thinking about what task you want to explore in your upcoming poster.
Use git to add, commit, and push¶
Be sure to save this notebook to the repository.